Rules
To participate in the challenge, please carefully review and comply with the following rules and requirements.
1. Challenge Structure
The competition includes one mandatory challenge and three optional challenges:
- Mandatory Challenge – Dengue (State Level): Forecast dengue cases at the state (UF) level for all Brazilian states, except Espírito Santo.
- Optional Challenge 1 – Dengue (City Level): Forecast dengue cases for 15 selected cities.
- Optional Challenge 2 – Chikungunya (State Level): Forecast chikungunya cases at the state (UF) level for all Brazilian states, except Espírito Santo.
- Optional Challenge 3 – Chikungunya (City Level): Forecast chikungunya cases for 10 selected cities.
2. Model Requirements
Your model must be hosted in a public repository on GitHub or GitLab.
The model must be registered on the Mosqlimate platform and named according to the following pattern:
3rd_imdc_{institution}_{team_name}- All characters must be lowercase.
institutionmust correspond to the acronym of the team leader’s institution.- In
team_name, you may choose any name you like, but it must contain only lowercase letters.
The repository must include a well-documented README to ensure reproducibility. The README must contain the following sections:
- Team and Contributors
- Repository Structure
- Libraries and Dependencies
- Data and Variables
- Model Training
- Data Usage Restriction
- Predictive Uncertainty
- References
You may submit more than one model. However, each submitted model must include predictions for all targets within the challenge(s) it is intended to participate in.
Additional details on how to complete the README and submit your model are available in the official template.
3. Data Usage Policy
- Any dataset not available via the FTP server or the Mosqlimate API must be submitted to the organizers.
- These datasets will be shared with all participants to ensure fairness and reproducibility.
4. Prediction Format and Requirements
4.1 Prediction Data Structure
Your prediction dataframe must include the following columns:
- date: Forecast target date (format: YYYY-MM-DD).
- pred: Median prediction (50th percentile).
- lower_x and upper_x: Prediction intervals of 50%, 80%, 90% and 95%. For example:
lower_95(2.5th percentile)upper_95(97.5th percentile)
These define a 95% prediction interval.
4.2 Validation Rules
To be accepted by the platform, predictions must strictly follow these rules:
- Sunday dates: All prediction dates must fall on Sundays (weekly resolution).
- Continuous dates: No gaps are allowed in the sequence of dates.
- Challenge timeframe: Predictions must cover all dates between Epidemiological Week (EW) 41 of the previous year and EW 40 of the target year.
- Positive values: All predictions must be greater than or equal to zero.
- Nested intervals: Prediction intervals must follow a consistent order:
lower_95 ≤ lower_90 ≤ lower_80 ≤ lower_50 ≤ pred ≤ upper_50 ≤ upper_80 ≤ upper_90 ≤ upper_95
Additional details on how to submit your predictions are available in the official template.
5. Submission Timeline and Forecast Targets
For each geographic unit (state or city, depending on the challenge), the following predictions must be submitted:
5.1 Validation Phase (Deadline: July 1)
Validation Test 1:
Forecast weekly cases for the 2022–2023 season (EW41 2022 – EW40 2023), using data from EW01 2010 to EW25 2022.Validation Test 2:
Forecast weekly cases for the 2023–2024 season (EW41 2023 – EW40 2024), using data from EW01 2010 to EW25 2023.Validation Test 3:
Forecast weekly cases for the 2024–2025 season (EW41 2024 – EW40 2025), using data from EW01 2010 to EW25 2024.Validation Test 4:
Forecast weekly cases for the 2025–2026 season (EW41 2025 – EW40 2026), using data from EW01 2010 to EW25 2025.
5.2 Forecast Phase
(After data update on July 31, deadline: September 10)
- Forecast Target:
Predict weekly dengue cases for the 2026–2027 season (EW41 2026 – EW40 2027), using all available data from EW01 2010 to EW25 2026.
6. Submission Completeness
- It is mandatory to submit both validation and forecast sets.
- Submissions must include predictions for all geographic units within each selected challenge.
- Incomplete submissions (missing geographic units or required prediction sets) will not be considered in the final evaluation.
7. Evaluation and Compliance
If any issues are identified in the model methodology or submitted predictions:
- The authors will be contacted and given the opportunity to make corrections.
- However, the organizers reserve the full right to exclude the model from the final reports submitted to the Ministry of Health if issues are not adequately resolved.
Please ensure full compliance with these guidelines to guarantee that your submission is valid and considered for evaluation. For further details, please refer to the Instructions tab.
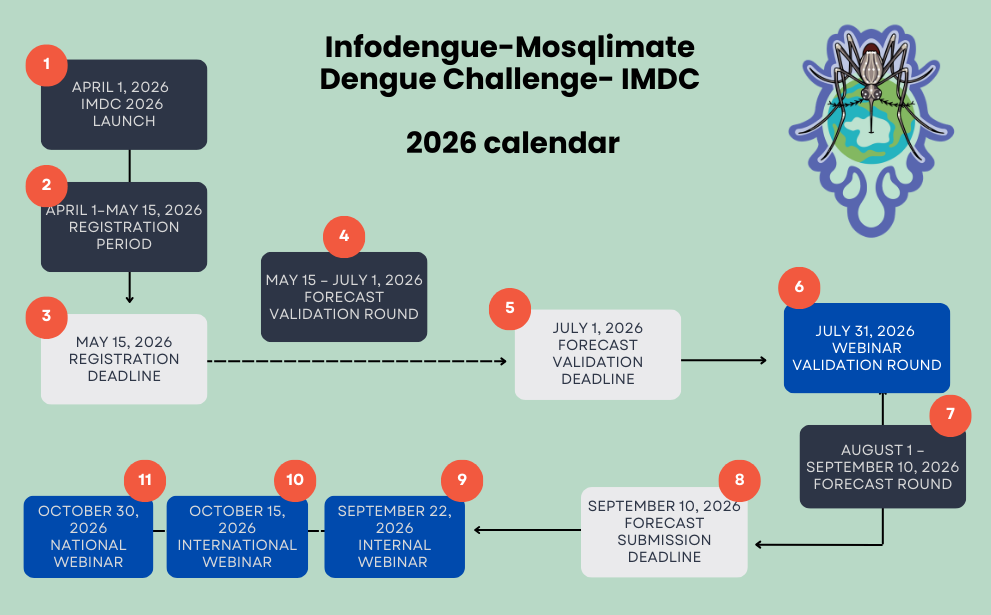
Important dates:
- April 1, 2026 – Challenge launch and opening of team registrations
- May 15, 2026 – Deadline for team registration
- July 1, 2026 – Submission deadline for validation round results
- July 31, 2026 – Webinar: Presentation of validation round results
- September 10, 2026 – Submission deadline for 2026–2027 forecasts
- September 22, 2026 – Internal webinar for teams to present model methodologies
- October 15, 2026 – International webinar: Technical results of IMDC 2026
- October 30, 2026 – International webinar: IMDC 2026 results for the general audience